Ecology of Freshwater Microbes
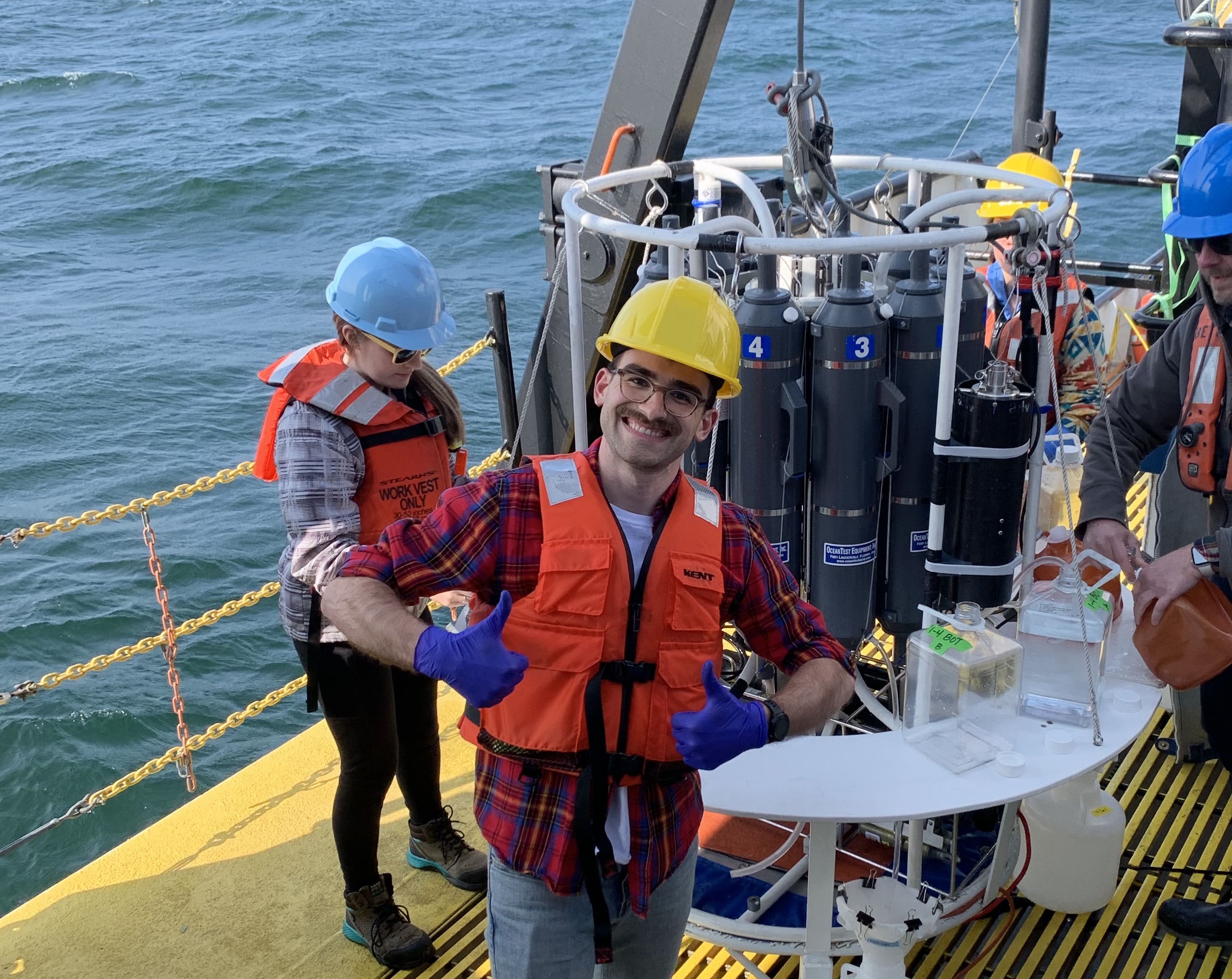
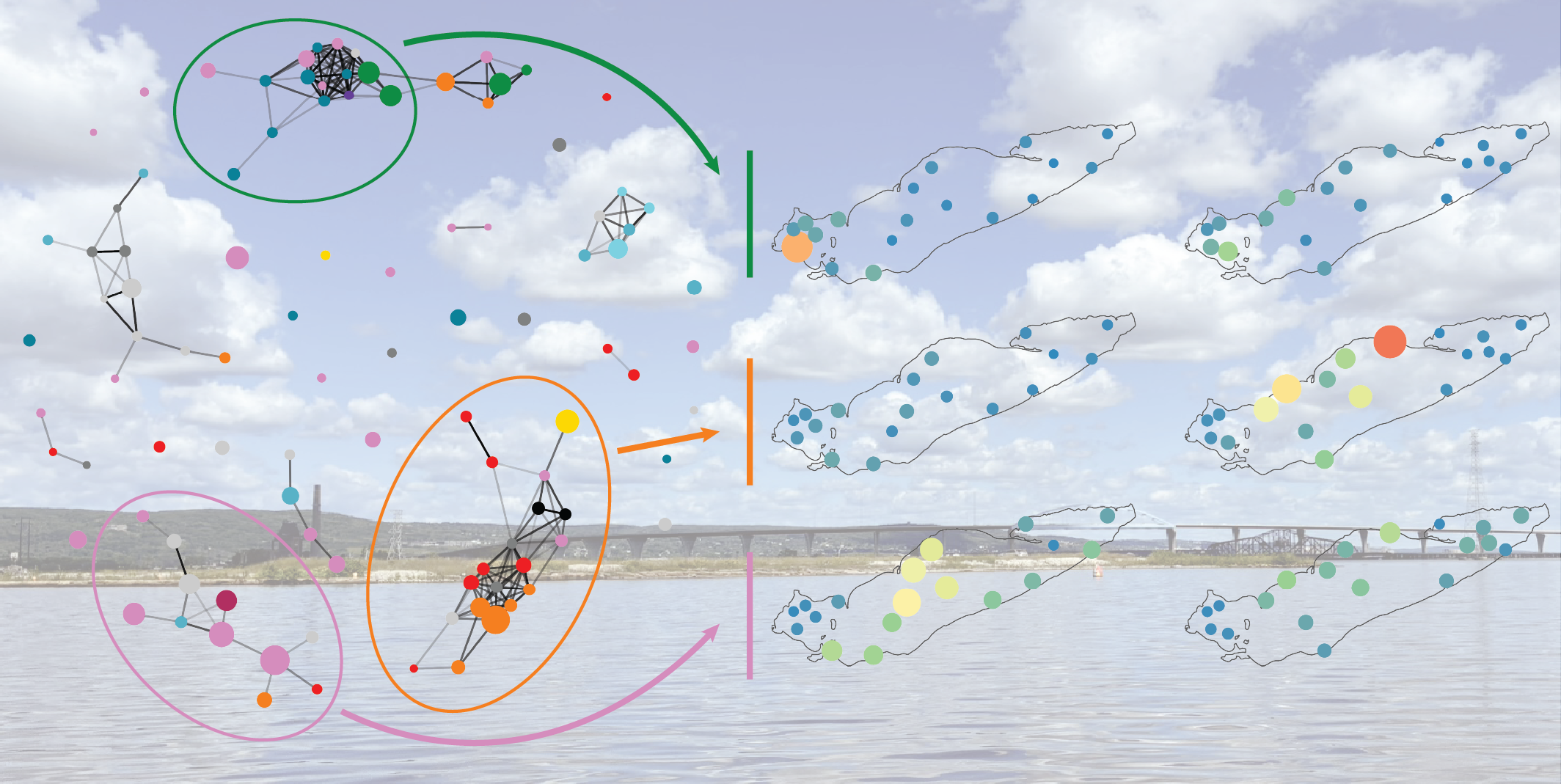
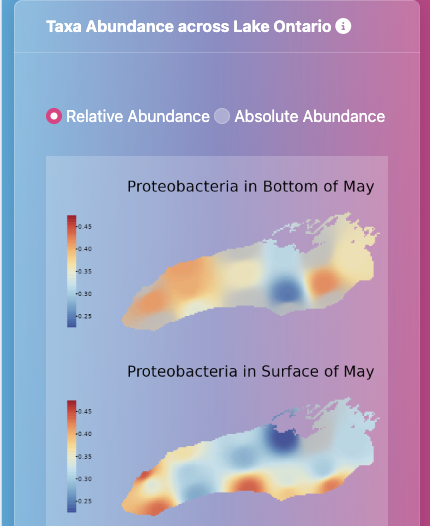
As a Ph.D. student in the Schmidt Lab, I study the formation, structure, and function of freshwater microbial communities in the Great Lakes. My fieldwork relies on collabortive relationships across EPA and NOAA NERR system, producing massive interdisciplinary datasets combining biological and environmental data types. I use bioinformatics, ecological modeling, and geospatial analysis to connect microbial diversity to hydrological processes. My work integrates both theory, as when we defined a new β-diversity metric, and empirical discoveries, as when we revealed how upwelling restructures Lake Ontario's microbiome.